Researchers have developed a new genome editing platform that allows precise manipulation of chromatin marks, revealing their direct impact on gene expression and challenging previous understanding of gene regulatory mechanisms.
A study by Hackett’s group at EMBL Rome has led to the development of a powerful gene editing technique, which opens up the ability to precisely program chromatin modifications.
Understanding how genes are regulated at the molecular level is a major challenge in modern biology. This complex mechanism is mainly driven by the interaction between proteins called transcription factors, DNA Regulatory regions and epigenetic modifications – chemical changes that alter the structure of chromatin. The collection of epigenetic modifications to a cell’s genome is referred to as the epigenome.
Advances in epigenome editing
In a study published today (May 9) in Nature geneticsScientists from Hackett’s group at the European Molecular Biology Laboratory (EMBL) in Rome have developed a modular genome editing platform – a system for programming epigenetic modifications anywhere in the genome. The system allows scientists to study the effect of each chromatin modification on transcription, the mechanism by which genes are transcribed into mRNA to catalyze protein synthesis.
Chromatin modifications are thought to contribute to the regulation of key biological processes such as growth, response to environmental signals, and disease.
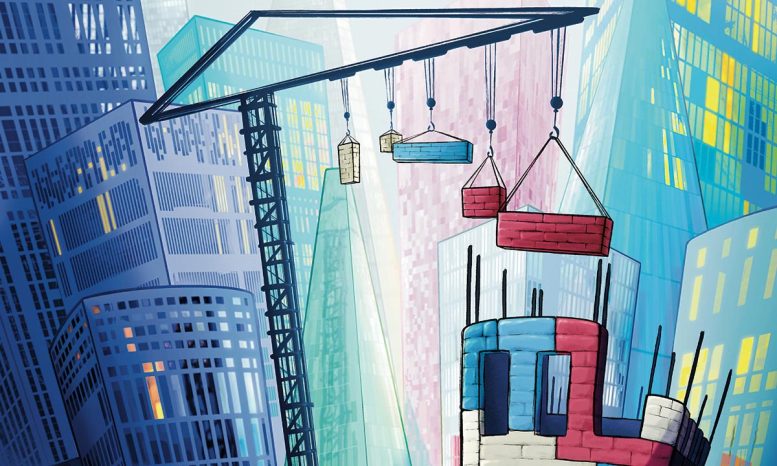

Creative illustration of the Epigenetic Editing Toolkit: Each building represents the epigenetic state of a single gene (dark windows are silent genes, light windows are active genes). The lever demonstrates the epigenetic editing system that enables de novo deposition of chromatin marks at any genomic site. Marzia Monafo
To understand the effects of specific chromatin marks on gene regulation, previous studies have mapped their distribution in the genomes of healthy and diseased cell types. By combining this data with gene expression analysis and the known effects of perturbing specific genes, scientists have attributed functions to these chromatin marks.
However, it has proven difficult to determine the causal relationship between chromatin marks and gene regulation. The challenge is to dissect the individual contributions of the many complex factors involved in such regulation – chromatin marks, transcription factors, and regulatory DNA sequences.
Breakthrough in epigenome editing technology
Scientists from Hackett’s group have developed a modular genome editing system to precisely program nine biologically important chromatin marks into any desired region of the genome. The system is based on CRISPR – a widely used genome editing technology that allows researchers to make modifications at specific sites of DNA with high precision and Accuracy.
Such subtle perturbations enabled them to carefully dissect cause-and-effect relationships between chromatin marks and their biological effects. The scientists also designed and used a “reporter system,” which allowed them to measure changes in gene expression at the single-cell level and understand how changes in DNA sequence affect the effect of each chromatin mark. Their results reveal the causal roles of a set of chromatin marks important in gene regulation.
Key findings and future directions
For example, researchers have found a new role for H3K4me3, a chromatin mark previously thought to be a consequence of transcription. They noted that H3K4me3 can actually increase transcription on its own if it is artificially added to specific sites in DNA.
“This was a very exciting and unexpected result that went against all of our expectations,” said Christina Policarpi, a postdoctoral researcher in Hackett’s group and lead scientist on the study. “Our data point to a complex regulatory network, where multiple governing factors interact to modulate gene expression levels in a given cell. These factors include the pre-existing structure of chromatin, underlying DNA sequence, and location in the genome.
Potential applications and future research
Hackett and his colleagues are currently exploring ways to harness this technology through a promising startup project. The next step will be to confirm and extend these conclusions by targeting genes across different cell types and on a broad scale. How chromatin marks affect transcription via gene diversity and downstream mechanisms remain to be elucidated.
“Our modular epigenetic editing toolkit constitutes a new experimental approach to dissecting the interrelationships between the genome and the epigenome,” said Jamie Hackett, group leader at EMBL Rome. “The system could be used in the future to more precisely understand the importance of epigenomic changes in influencing gene activity during development and in human diseases. On the other hand, this technology also opens up the ability to program desired gene expression levels in a highly tunable way. This is an exciting avenue for applications.” Accurate health benefits may be useful in cases of illness.
Reference: “Stem genome editing captures the context-dependent instructive function of chromatin modifications” 9 May 2024, Nature genetics.
doi: 10.1038/s41588-024-01706-s

“Explorer. Unapologetic entrepreneur. Alcohol fanatic. Certified writer. Wannabe tv evangelist. Twitter fanatic. Student. Web scholar. Travel buff.”